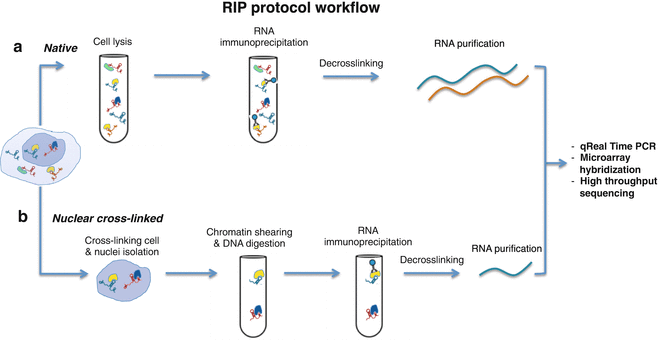
Figures and data in The unfolded protein response and endoplasmic reticulum protein targeting machineries converge on the stress sensor IRE1 | eLife
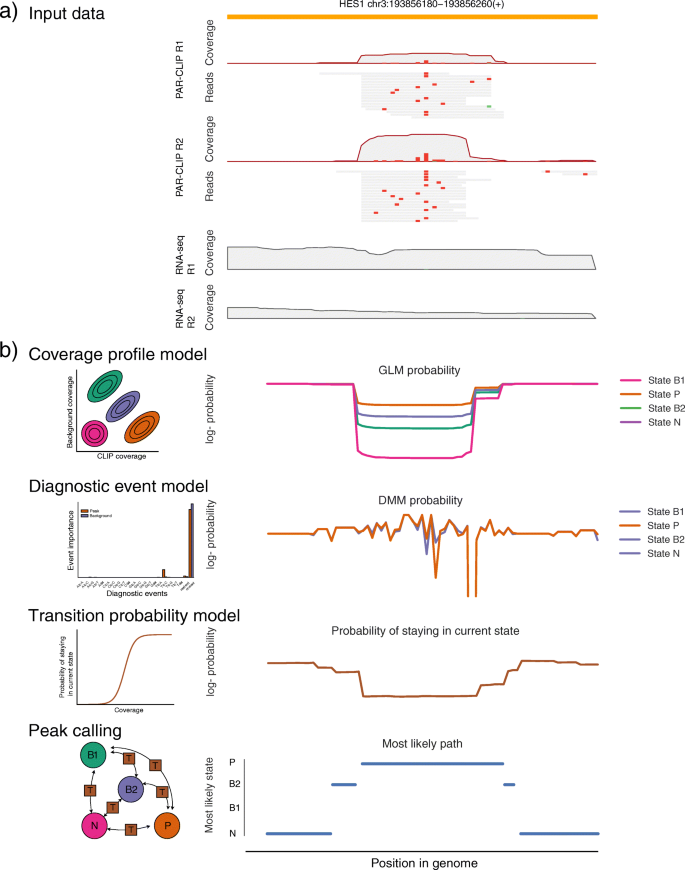
omniCLIP: probabilistic identification of protein-RNA interactions from CLIP-seq data | Genome Biology | Full Text

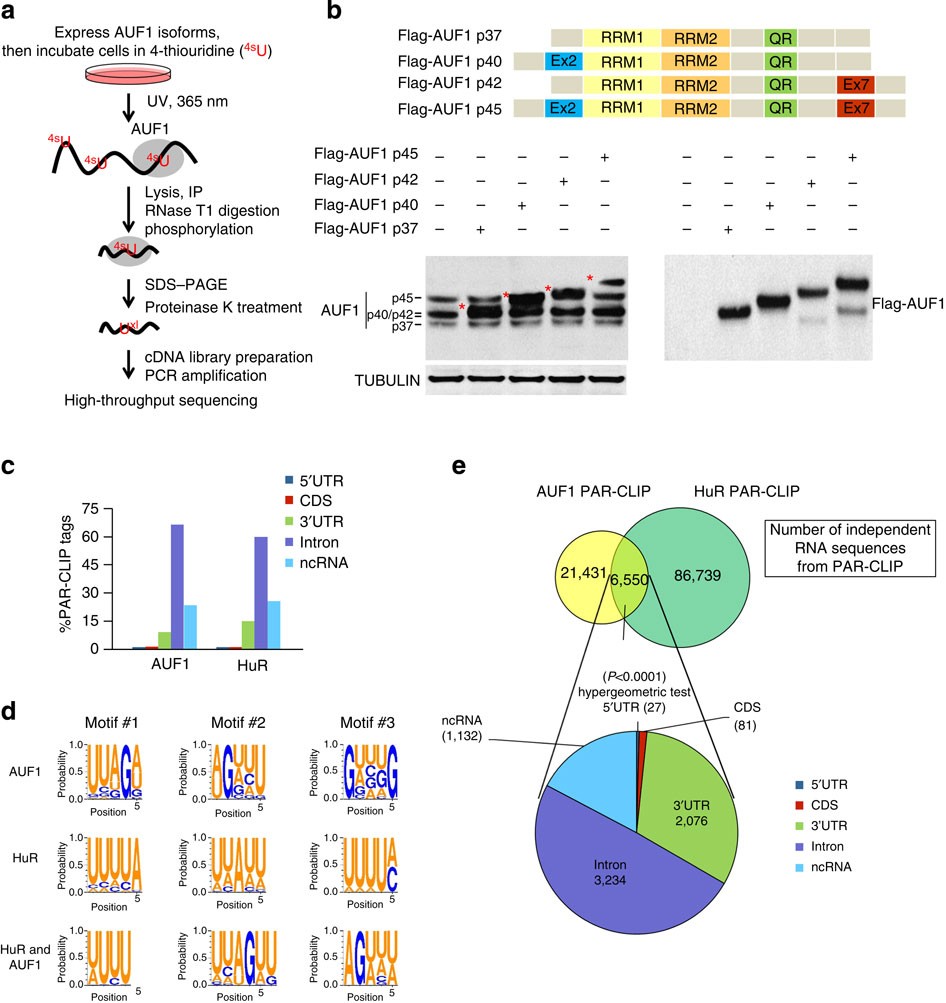
Transcriptome-wide Identification of RNA-Binding Protein and MicroRNA Target Sites by PAR-CLIP: Cell

Global Approaches in Studying RNA-Binding Protein Interaction Networks: Trends in Biochemical Sciences
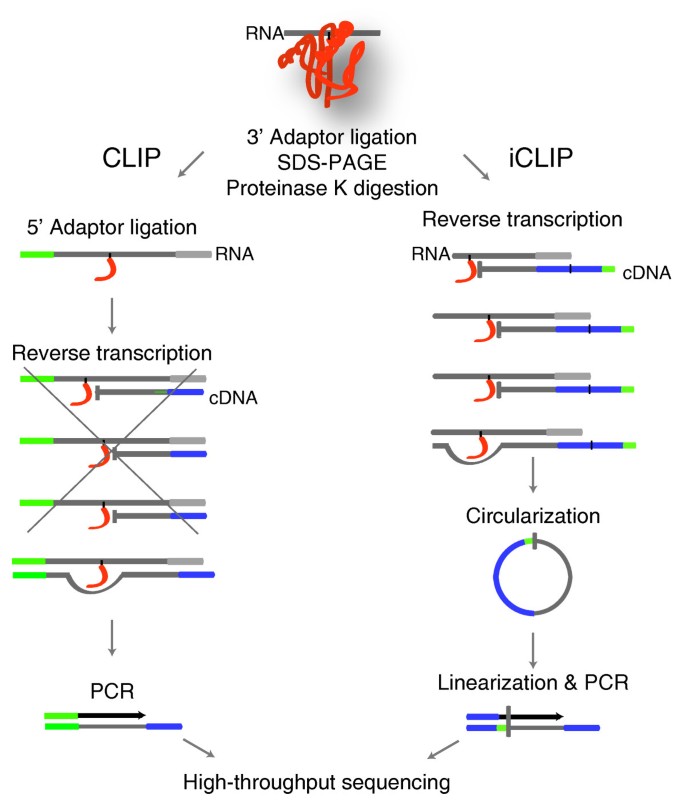
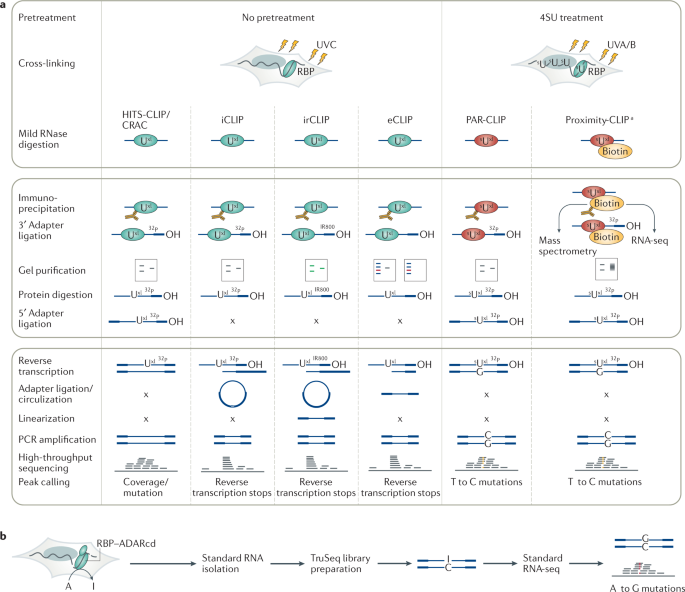
Analysis of CLIP and iCLIP methods for nucleotide-resolution studies of protein-RNA interactions | Genome Biology | Full Text

Transcriptome-wide Identification of RNA-Binding Protein and MicroRNA Target Sites by PAR-CLIP - ScienceDirect

PAR-CLIP and streamlined small RNA cDNA library preparation protocol for the identification of RNA binding protein target sites - ScienceDirect

Optimization of PAR-CLIP for transcriptome-wide identification of binding sites of RNA-binding proteins - ScienceDirect
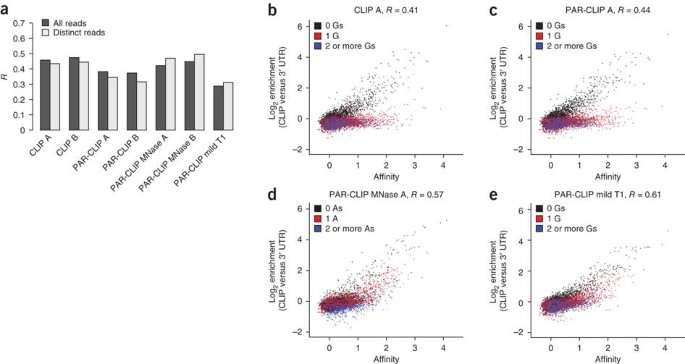
A quantitative analysis of CLIP methods for identifying binding sites of RNA-binding proteins | Nature Methods

Transcriptome-wide Identification of RNA-Binding Protein and MicroRNA Target Sites by PAR-CLIP: Cell

PAR-CLIP: A Method for Transcriptome-Wide Identification of RNA Binding Protein Interaction Sites | SpringerLink

Outline of HITS-CLIP, PAR-CLIP and several variants, iCLIP, iCLAP and... | Download Scientific Diagram





![PDF] Current methods in the analysis of CLIP-Seq data | Semantic Scholar PDF] Current methods in the analysis of CLIP-Seq data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/e504d75c2c48b19613a796fb54083b71e3781289/2-Figure1-1.png)